
Database acceleration using specialized co-processors is nothing new. Just to give a few examples, data warehouses running on the Netezza platform, owned by IBM for more than a decade now, uses a custom and parallelized PostgreSQL database matched to FPGA acceleration for database and storage routines. OmniSci, Sqream Technologies, Kinetica, and Brytlyt all have GPU acceleration for their analytics databases, and have seen moderate success goosing performance for gigascale and even terascale databases with up to tens of billions of rows.
But TigerGraph, the upstart maker of the graph database by the same name, has been working with FPGA maker Xilinx on a new twist on database acceleration and is putting a stake in the ground right here at The Next Platform to deliver petascale graph databases — a tall order because graph databases are notoriously hard to scale — within the next two years.
The person driving that stake into the ground for TigerGraph is Jay Yu, vice president of product and innovation, who just joined TigerGraph in August being a distinguished engineer, architect, and director at financial application services company Intuit for nearly 19 years. Yu got his PhD in computer science and database systems at the University of Wisconsin, and studied under David DeWitt, a luminary in the database world who is now at MIT. While at the university, Yu worked on the Paradise shared nothing, object relational database management system, which was acquired by Teradata primarily for its distributed query execution engine.
While at Intuit last year, Yu led the team that was doing a bake-off of databases to be used for terabyte-range graphs that would underpin fraud detection, entity resolution, and so-called “customer 360” databases that are used by the Intuit software stack to serve the 100 million customers worldwide that use its TurboTax, QuickBooks, Mint, and Credit Karma online applications. TigerGraph won the bakeoff, by the way, and not only did Yu discover that some of the distributed computing ideas embodied in the Paradise database had been used in TigerGraph, the company offered Yu a job to come help scale the product and push it out into the world.
For those of you unfamiliar with TigerGraph, we did a detailed story on the company when it dropped out of stealth three years ago, and during our The Next Database Platform event in September 2020, we interviewed Yu Xu, founder and chief executive officer at TigerGraph, as well as with Edward Sverdlin, vice president of advanced technology, research, and development at UnitedHealth Group, about building the world’s largest known healthcare graph.
“We have already run LDBC benchmark at the 36 TB level, and other vendors are struggling at the 1 TB level, explains Yu to The Next Platform, referring to the Linked Data Benchmark Council Social Network Benchmark test, which has two workloads — one interactive that does transactional graph processing on a node and its neighbors, inserting data, and the other a business intelligence workload that has a lot of aggregations and joins on very heavy queries that touch a large portion of the graph. “And that number was not even with hardware acceleration. We are working on a 100 TB version of the LDBC-SNB, and you probably need to give us one or two quarters to do that. And in one or two years, we want to reach a 1 PB graph. And I am so excited to push this forward because if TigerGraph can handle petabyte-level graphs, I will argue that the data lakes at most companies should just be stored in TigerGraph abd it is already preorganized.”
As we have said before, the beauty of a graph database is that it has its index built in by default and you don’t have to do joins anymore.
Hardware acceleration is going to be a big part of getting TigerGraph up to petascale. So here is how it works.
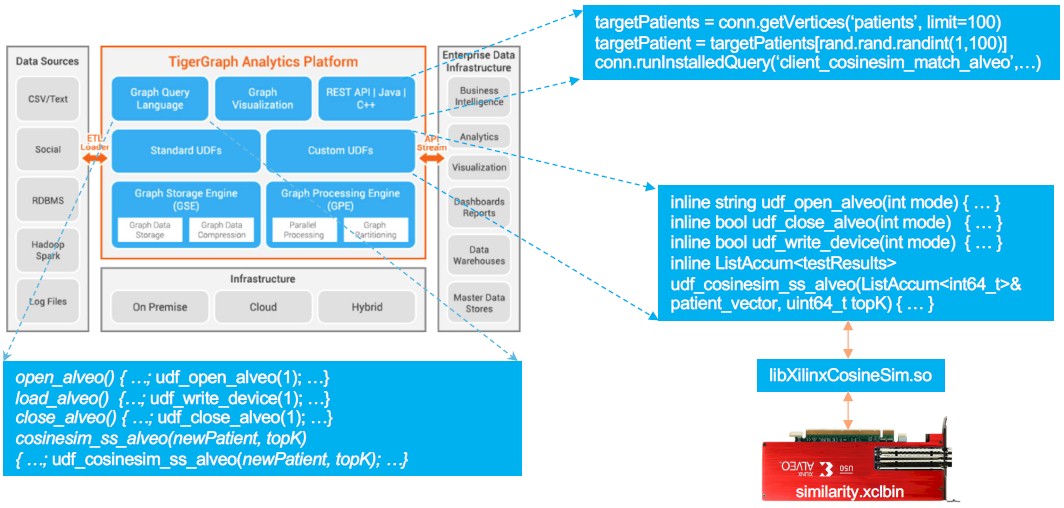
Within the TigerGraph database, says Yu, there are about 50 different algorithms that do the processing for the graph; about twelve of them are core to the graph database, meaning they are either heavily used, or have heavy computational loads, or both. To test out the acceleration idea, TigerGraph worked with Xilinx to take two such routines — cosine similarity and the Louvain method of modularity optimization — and implement these algorithms in the programmable logic of the FPGA, where they are offloaded from the CPU. Conceptually, here is how TigerGraph and the Xilinx Alveo U50 FPGA coprocessors were integrated:
The Xilinx FPGAs are running a modified Vitis graph analytics library that that does the cosine similarity and top 1,000 scores for similarities — in this case, the workload was looking at healthcare patient records stored in a graph database for similar cases.
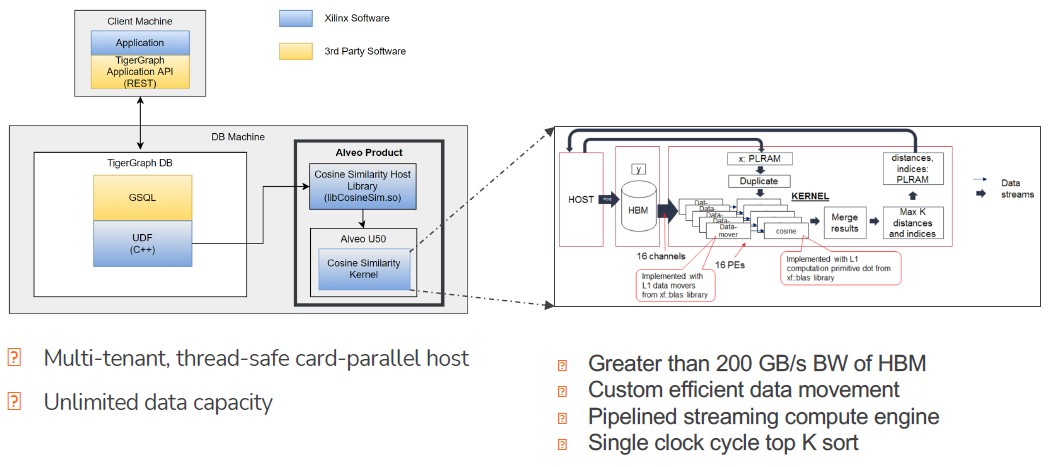
Here is a better schematic for the cosine similarity workload:
The GSQL queries are compiled down to user defined functions in C++ by the Vitis stack and then passed off to the cosine similarity functions in the Vitis library and run on the Alveo U50s. There are multiple cards that can be used to accelerate the workload, and these cards are run in parallel fashion now, but eventually there will be interconnects — probably CCIX interconnects and maybe someday after the AMD acquisition of Xilinx is done, Infinity Fabric links — between the FPGAs so they can share data directly, like Nvidia does with NVLink ports and its NVSwitch switch for its GPU accelerators.
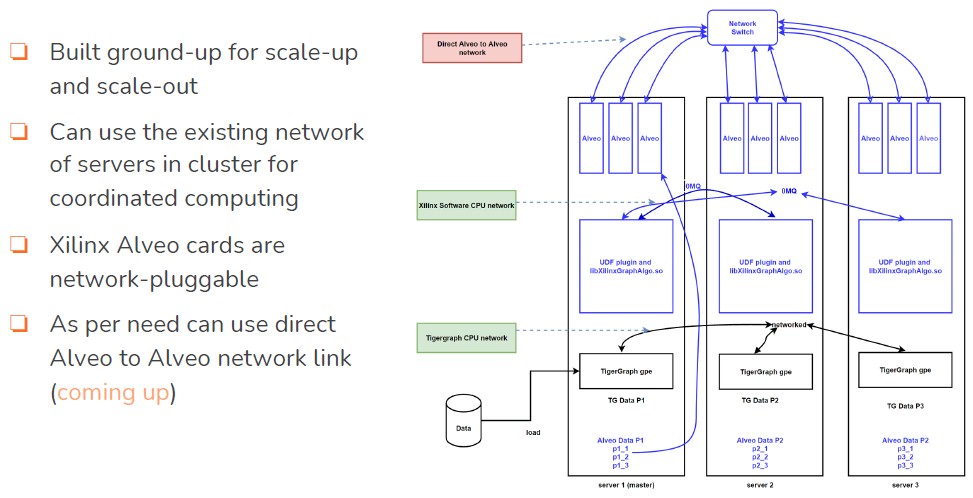
Here is what the database cluster architecture looks like:
Incidentally, the FPGA acceleration is not so much about what we call scale in, which means doing more work within a single node to reduce the size of a cluster, and it is not so much about scale out, which means using multiple nodes to get an increasing amount of data stored and usually an increasing amount of work done as the node counts go up. This latter one is in contrast with scale up, which means getting a single node to scale up its address space and compute to run a workload that has not been tuned for cheaper scale out architectures. The FPGA acceleration that TigerGraph is doing is about reducing the amount of time for graph calculations, which will not cut down the size of a TigerGraph cluster. But this will reduce the time to do certain kinds of queries and calculations, which increases the number of iterations an organization can do as they sift through their data for connections. The performance increases are substantial, as you will see in a moment.
The benchmark tests that TigerGraph and Xilinx did were performed on ProLiant DL385 Gen10 Plus servers from Hewlett Packard Enterprise, which have a pair of AMD “Milan” Epyc 7713 processors with 64 cores running at 2 GHz. This server had 512 GB of main memory and 60 TB of flash storage, a mix of SATA and NVM-Express devices. The machine was equipped with five Xilinx Alveo U50 cards, which cost $2,500 a pop by the way (reasonably priced compared to some GPU accelerators in the datacenter). The Alveo U50 cards, which we covered here, have 6,000 DSP blocks, 872,000 lookup tables (LUTs), 1.7 million flip-flops, 227 MB of on-chip block RAM, and 8 GB of HBM2 memory with 460 GB/sec of bandwidth; they burn under 75 watts. The system ran Ubuntu Server 20.04.1 LTS and had TigerGraph 3.1 Enterprise Edition loaded along with the Vitis libraries. The test was run against the Synthea synthetic patient dataset.
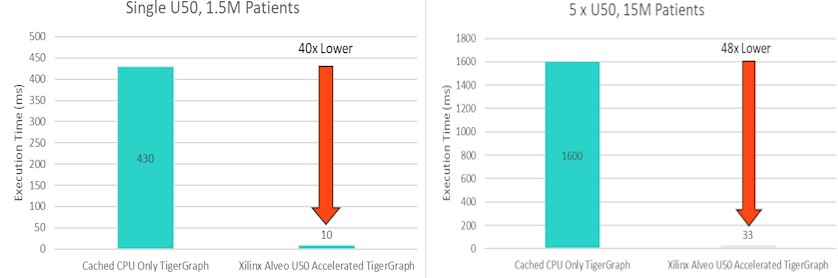
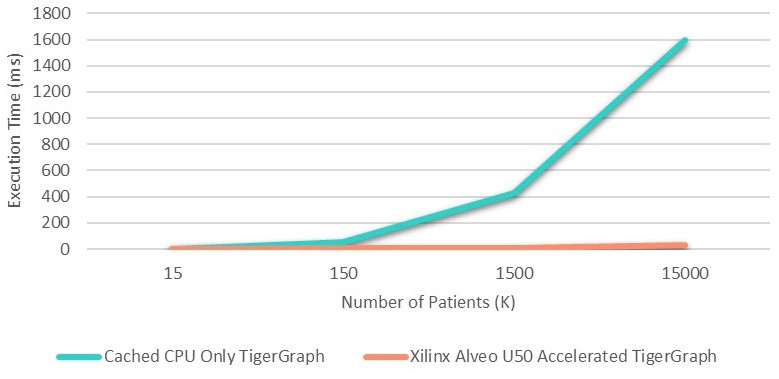
On a subset of the data with 1.5 million patients and one FPGA turned on and 15 million patients with five FPGAs turned on, here is what the CPU-only and Alveo-boosted response times looked like trying to find the top 1,000 similar patients with the cosine similarity function accelerated on the FPGAs:
The CPU-only response time was 430 milliseconds on average to find 1,000 most-alike patients in the 1.5 million patient dataset, and it dropped down to 10 milliseconds with the FPGA accelerating the cosine similarity function. Increasing the dataset size by a factor of 10X to 15 million patients, the CPU-only response time was 1.6 seconds — not great — but was 48X lower at 33 milliseconds with five FPGAs activated.
The response time goes up pretty fast as the CPU-only scenario struggles to do all of the math, as you can see in the chart below, but the FPGA accelerated cosine similarity function really helps to flatten response times:
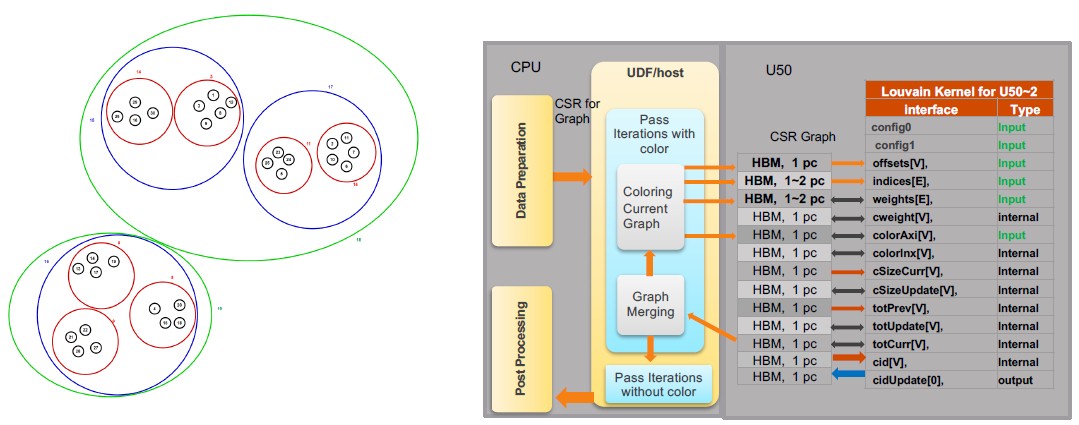
With the Louvain modularity algorithm, the graph database is scanned to find community grouping across entities. Like this:
According to Yu, the Louvain modularity method is one of the most CPU-intensive algorithms in graph databases, and it is not a deterministic one but a probabilistic one. And the more CPU and memory you throw at this algorithm, the more accurate it becomes. In this case, the FPGA acceleration provided a 20X boost in response time, but also the accuracy of the groupings of people was 18 percent more accurate. (We didn’t have charts for this one. This is a big deal if you can move from 20 minutes to 1 minute, or from 20 seconds down to 1 second to do these groupings.)
The neat thing is that Xilinx FPGAs are available on many of the big public clouds, TigerGraph runs on cloud instances there, and companies can test the whole shebang out before building up their own infrastructure.
When the dozen functions are ported to FPGA acceleration, TigerGraph’s performance should improve over many functions, and when and if all 50 algorithms are accelerated, this will be a big help. And maybe by that time, CPUs will have some of these functions encoded in their transistors, making it all even easier. There are lots of possibilities. By the way, none of this work is tied to Xilinx FPGAs — it will work with Intel FPGAs, too, says Yu. And some functions could also be accelerated by GPUs if that is the accelerator of choice, too. But TigerGraph is a startup and it has to focus on the easiest things with the fastest returns or the most important things with the biggest returns. Like petabyte-scale graph databases, for instance.





Be the first to comment