
A collaboration of researchers from the University of California Davis, the National Energy Research Scientific Computing Center, and Intel are working together on the DisCo project to extract insight from complex unlabeled data.
DisCo is short for the Discovery of Coherent Structures, and it discovers the inherent structures in unlabeled data, which, given the nature of high-dimensional data, requires innovative clustering methods and a powerful system like the Cori supercomputer at NERSC. The discovered structures are then used in subsequent analysis to provide much greater detail beyond simple summary statistics traditionally used when studying high-dimensional systems.
According to Karthik Kashinath, project scientist at NERSC: “The DisCo project discovers patterns in spatiotemporal data in an unsupervised fashion. This addresses a major challenge of deep learning, a form of supervised machine learning that requires large amounts of labeled training data.”
A paper, DisCo: Physics-based Unsupervised Discovery of Coherent Structures in Spatiotemporal Systems was presented at the 2019 IEEE/ACM Workshop on Machine Learning in High Performance Computing Environments (MLHPC).
Adam Rupe, a physics PhD candidate at UC Davis, emphasizes that “DisCo is a principled, physics-based tool that scientists can use to discover and identify the key components that organize macroscopic behavior in complex spatiotemporal systems, like extreme weather events in atmospheric dynamics.” This is achieved using mathematical objects known as local causal states that draw upon the physics of self-organization to extract pattern and structure directly from a system’s dynamical behavior, without requiring the governing equations or pre-specified templates or labeled training examples.
Python for Prototype And Production
A big achievement of the DisCo project is successfully using Python for both prototyping the machine learning pipeline as well as deploying at scale in a production HPC environment. Python is extremely popular amongst domain science researchers and data scientists.
Typically, these Python prototypes must be rewritten for production deployment using lower-level languages like C/C++. While this produces high-performance code, it can incur considerable costs developer productivity. The ability to run Python code in production by achieving similar performance as the C/C++ code is important. This was brought to fruition by using Intel Distribution of Python which is optimized for Intel platforms, featuring highly-optimized libraries such as Intel MKL and Intel DAAL. Members of the DisCo team (from Intel) added support for distributed clustering via Intel daal4py and optimized library functions for the high-dimensional problem datasets used in DisCo. These and other open source libraries and tools such as Numpy and the Numba open source JIT compiler allowed the team to optimize the full pipeline. By parallelizing and vectorizing portions of the Python pipeline that are not part of Intel MKL and Intel DAAL, they ensured the end-to-end machine learning workflow was optimized for performance.
“We use Python for both prototype development and production deployment on an HPC system”, notes Nalini Kumar (engineer, Intel), “This brings us closer to bridging the performance and productivity disconnect, and streamlining the process from theoretical development to deployment at scale for domain applications.”
An HPC First
Project DisCo combines the first distributed HPC implementation of local causal state reconstruction with theoretical advances in using local causal states to decompose spatiotemporal systems into structurally relevant components.
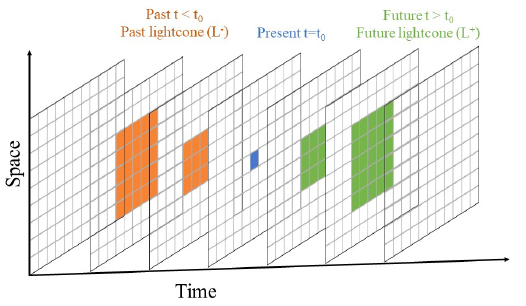
Local causal states are reconstructed using a notion of “causal equivalence” between points in spacetime, based on their past and future lightcones (orange and green features, respectively, in the figure below). The DisCo algorithm approximates causal equivalence using two stages of clustering, assigning the same local causal state label to all effectively equivalent points. The reconstructed local causal states then identify distinct space-time structures in the data.
Core to the DisCo pipeline is the first clustering stage that performs distributed distance-based clustering over the high-dimensional lightcones (potentially over one thousand dimensions). The team worked with two well-known clustering algorithms – K-Means and DBSCAN (Density-based spatial clustering of applications with noise) – to cluster lightcones.
The DisCo team used sample segmentations images for a 2D turbulence simulation, videos from the NASA Cassini Jupiter mission, and water vapor flows from a CAM5.1 climate model simulation to test the spacetime segmentation.
The data can be quite complex as illustrated in the following snapshot images taken from videos. The colored regions constitute a lightcone, or representation of the spatial data through time. DisCo refers to a speed-of-light value, c, to represent how fast information propagates temporally.
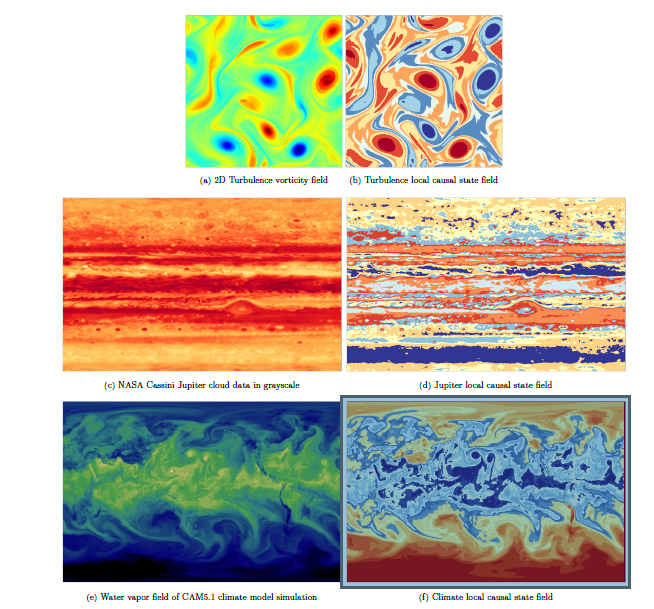

Images in the left column are snapshots from the unlabeled “training” data used for local causal state reconstruction. Images in the right column are corresponding snapshots from the local causal state segmentation fields.
As can be seen in the right column, the DisCo results appear to segment (discover space-time structures) quite nicely from the input data. These results, the team states, “demonstrate state-of-the-art segmentation results for three complex scientific datasets, Including on 89.5 TB of simulated climate data (lightcones) in 6.6 minutes end-to-end.” The full segmentation videos are available at the DisCo YouTube channel.
Both K-Means and DBSCAN weak-scaling and strong scaling experiments were run on the CAM5.1 climate dataset. K-Means experiments were run on Cori’s “Haswell” Intel Xeon E5 processors and DBSCAN experiments were run on both Cori’s Xeon E5 and “Knights Landing” Xeon Phi processors.
Hero Run With K-Means
Evaluation of small datasets proved K-Means clustering was superior to DBSCAN for the problems evaluated. Thus, the team chose the distributed K-Means clustering method for their hero run. During this run they processed 89.5TB of lightcone data, obtained from 580 GB of CAM5.1 climate simulation data, on 1024 Intel Xeon nodes with 2 MPI ranks/node and 32 tbb threads/processor.
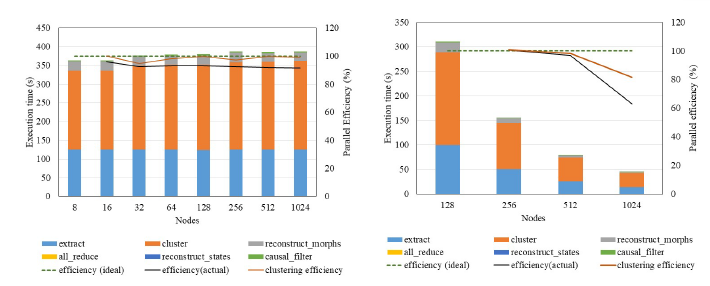
The total time to solution was 6.6 minutes with a weak scaling efficiency of 91 percent. In the figure below, the left column shows weak scaling results (91 percent scaling efficiency) and the right column shows the strong scaling results (64 percent scaling efficiency). Results from the hero run and the weak scaling experiments exhibit that the DisCo code scales well to process huge data sets and enable physics-based discovery in realistic systems.
Weak scaling and strong scaling DBSCAN was performed using the climate data set on both Intel Xeon and Intel Xeon Phi nodes.
For weak scaling on Intel Xeon and Intel Xeon Phi nodes, the spatial field was split across eight nodes. Each local node clustered its data set before merging with different nodes. A scaling efficiency of 34.6% was observed. For DBSCAN, the total running time is dictated by the runtime of the slowest node. Load imbalance in the number of points per node is a contributing factor.
For strong scaling the team used a 1152 x 768 spatial field per node for the 128-nodes run; one timestep across 2 nodes for the 256 node run; one timestep across 4 nodes for the 512 node run; and one timestep across 8 nodes for the 1,024 node run. Overall scaling efficiency was 38 percent on Xeon Phi nodes and 52 percent on Xeon nodes.
Open Source And Vendor Optimized Libraries Are Essential
The team further notes that achieving Python performance and scalability on HPC systems relies heavily on availability of highly-optimized vendor libraries with easy-to-use APIs and integrated seamless distributed memory processing modes with popular libraries like scikit-learn, JIT compilers like Numba, and more.
The heavy reliance on both open-source and vendor optimized libraries can be seen in the following list from the paper, “We use Intel Distribution of Python (IDP) 3.6.8. IDP incorporates optimized libraries such as Intel MKL and Intel DAAL for machine learning and data analytics operations to improve performance on Intel platforms. We also use NumPy (1.16.1), SciPy (1.2.0), Numba (0.42.1), Intel TBB (2019.4), Intel DAAL (2019.3) and Intel MKL (2019.3) from Intel Anaconda channels. MPI4Py (3.0.0) is built to use the Cray MPI libraries. All K-Means experiments used Intel Daal4py (0.2019.20190404.223947) built from source.”
Validating unlabeled results that are otherwise not externally well-defined is a challenge faced by any unsupervised method. Like with extreme weather events, there is no consensus on how to properly define more general structures in complex fluid flows. Ground truth for these structures does not currently exist.
This meant that, for the more complex climate and Jupiter datasets visual inspection of the output from DisCo was used to determine whether the “structural” decomposition of the local causal states capture meaningful “structure” in the flow. However, the team did compare the 2D turbulence and Jupiter results with current state-of-art using Lagrangian Coherent Structure methods, but that is beyond the scope of this article (see the paper for more detail).
In looking to the future, DisCo researchers make the following claims in their paper:
- First distributed-memory implementation of local causal state reconstruction opening the door to processing very large real-world causal data sets.
- Performs unsupervised coherent structure segmentation on complex realistic fluid flows.
- Achieved good single-node, weak scaling, and strong scaling performance up to 1024 nodes.
- Use Python to achieve high performance while maintaining developer productivity by using a vendor optimized version of Python, various supporting libraries, and compilers.
Rob Farber is a global technology consultant and author with an extensive background in HPC and in developing machine learning technology that he applies at national labs and commercial organizations. Rob can be reached at info@techenablement.com.





Be the first to comment