
Hard to believe, but the R programming language has been with us since 1993.
A quarter century has now passed since the authors Gentleman and Ihaka originally conceived the R platform as an implementation of the S programming language.
Continuous global software development has taken the original concepts originally inspired by John Chambers’ Scheme in 1975 to now include parallel computing, bioinformatics, social science and more recently complex AI and deep learning methods. Layers have been built on top of layers and today’s R looks nothing like 1990’s R.
So where are we at, especially with the emerging opportunities for deep learning on the horizon?
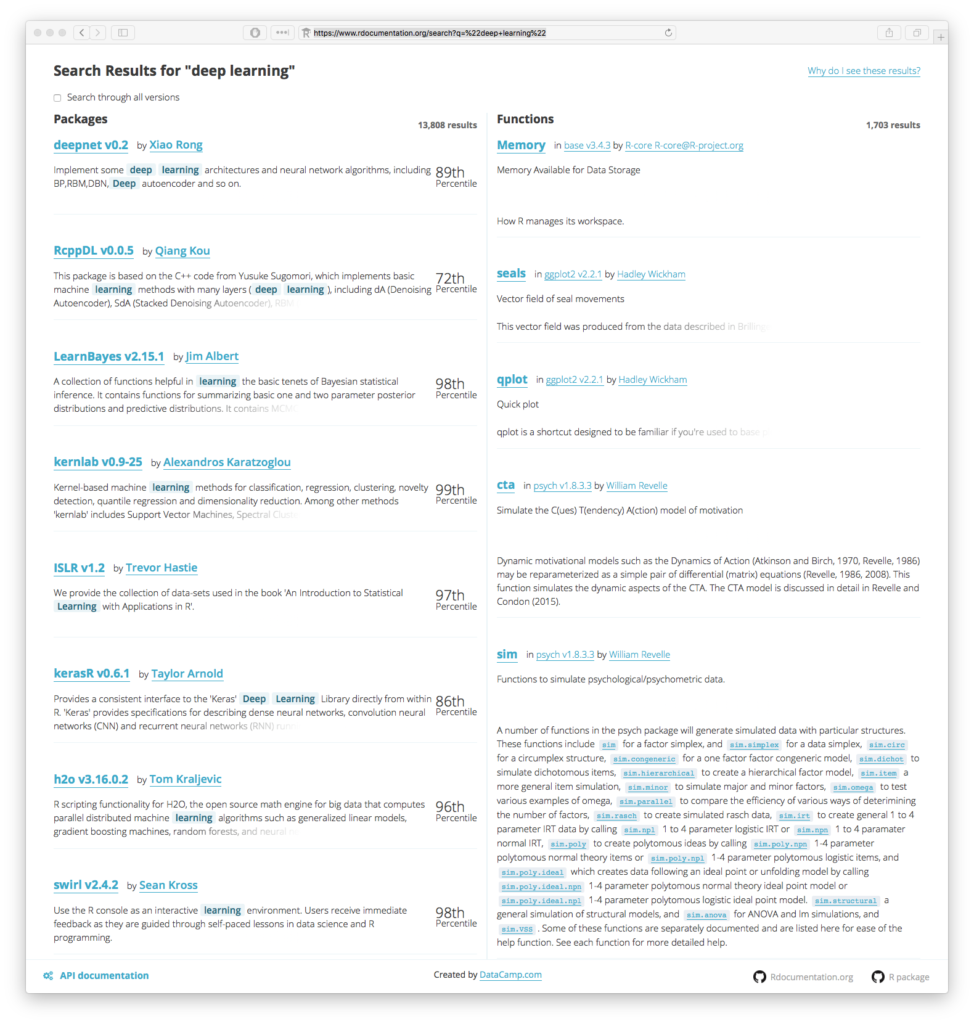
Current state-of-the-art leadership for deep learning in R is provided by a whole slew of “bolt on” packages, algorithms and methods. Keras, H20.ai and mxNet are currently some of the more visible and well executed methods for integration with R. Each of these three round out an impressive array of other independent development sitting on top of the core R stack. Each are attempting to provide what we here at The Next Platform have been calling “Easy AI”. It is a full on popularity contest for sure. Thousands of results are returned from searching the Rdocumentation project for the keywords “Deep Learning” alone.
Some of the most commonly asked questions today include “Which package should I use for Deep Learning? What’s the best Deep Learning method?”. It feels like every data scientist has been given some sort of homework assignment by their leadership. Each of them are desperately rooting about aimlessly in the dark recesses of the internet in a vain attempt to get a quick answer before their homework is due to teacher on Monday morning. It’s not right.
Our own research here at The Next Platform showed a number of the usual suspects bubble to the top of any list. But the challenge of answering which are the “best”? Well that is becoming an increasingly more difficult and potentially impossible question to answer. The very fact that there are over 14,500 total packages to choose from in CRAN, Github and Bioconductor tells you how complicated and fragmented development with R is.
For each of these packages, multiple versions also exist, couple that with the fact that a new version of the R core software is released every few months, scientific provenance and reproducibility here is the real challenge. Sure you can just stick it all in a container, but there’s way more to it than that.
The deep learning popularity contest continues, it’s not just how many times a package has been downloaded, but how efficient it is, can it exploit GPU or parallel learning, does it continue to give the same answer after an upgrade? These are the real questions to ask. The only way we can answer these questions effectively is we need to select methods that most efficiently and accurately document themselves.
So, taking just the top few methods, mxnet, darch, deepnet, h2o, and keras as an example will give you potentially hundreds of all-by-all comparisons to be made to make your selection. The size and scope of algorithm selection and execution is clearly outside even a The Next Platform long form article and is now becoming the basis of multiple research studies in and of themselves, and in-depth online tutorials. So let’s assume you do have to pick only one. What do we suggest you pick?
Documentation must drive selection
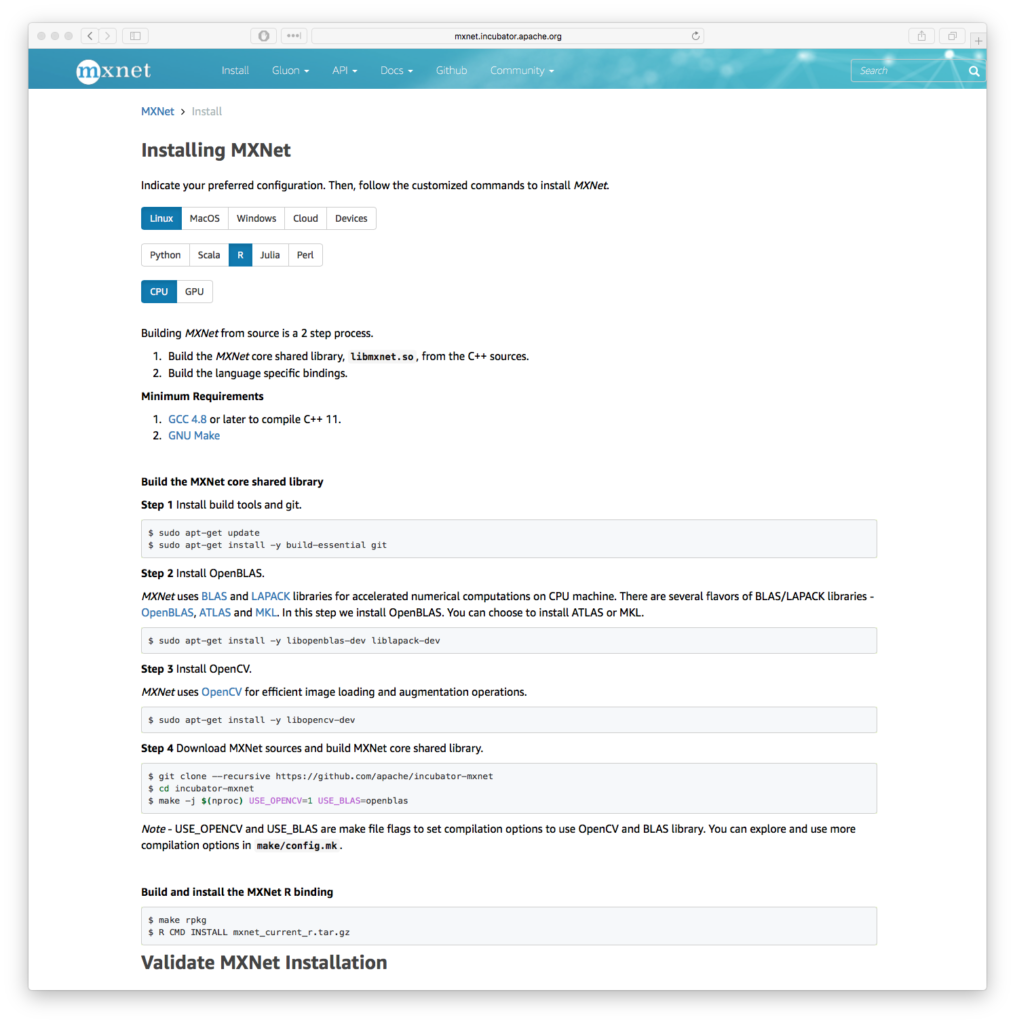
Many, many packages exist with scant or obscure documentation reflecting the “science project” nature of partially complete development processes are unfortunately very common with a number of R packages. However, projects such as MXNet clearly stand out. Succinct “getting started” and “how to” guides are key in this new world order of package variety and complexity. Gluon is a further attempt by the project to provide simple interfaces into the complexity. You can quickly see that time has been taken to clearly document the system and interfaces, which is a hallmark of a number of other Apache Foundation software products. Simple guides to provide rapid access to training modules is critical here.
For example, there are 12 options to install MXNet, including R as one of the platforms including a few prerequisites like our old favorites BLAS, LAPACK and chums. It’s a whole lot of stuff to build, you need good examples and documentation. The Next Platform was able to quickly reproduce MXNet installation and example code while researching this article on their own systems because the documentation was both clear and accurate. For comparison, older deep learning toolkit documentation was a little more terse, you can quickly see how we have developed better methods to document complex software.
As a result of library and operating system complexity is also the reason why more and more researchers are asking “for root” on shared systems. Installing the vast number of components and algorithm dependencies are really driving the need for systems such as Singularity and Singularity-Hub and others to somehow “contain” these ever increasingly complex stacks. Even executing one of these “Deep Learning in R” processes relies on millions upon millions of individual system calls to even load all the required pieces in to memory. That process all has to happen before you even start running your own workloads.
The challenge here is being taken on by commercial ventures. Being able to package systems and make “Easy AI” happen is absolutely non trivial. H2O.ai now claim over 129,000 data scientists in over 12,000 organizations are using their stack. However, they also quickly realized that crystal clear documentation is key, as is solving for the challenge of “abstraction”. By providing commercial support and detailed documentation H2O.ai can help organizations get on board with a complex set of software and systems as quickly as possible. Recent efforts around their “Driverless AI” product set further abstracts the complexity from the end user of dealing with 1,000’s of little bits and pieces of software.
So what about back to rolling your own? Tensorflow for R ends up presenting itself via a Keras API. To be clear this is R on top of python on top of Tensorflow. It’s a deep stack. However, the recently released book “Deep Learning with R” by Francois Chollet with J. J. Allaire helps to unravel the confusion. Chollet is the creator of Keras and a researcher within the deep learning group at Google, Allaire cofounded RStudio and authored the R interfaces into TensorFlow and Keras. Between them, they have seen and experienced this topic at the very sharp end. Deep Learning with R will no doubt be the future reference work on how R and Deep Learning techniques can hang out with each other and make beautiful music together. Nothing documents a system better than a detailed and long form book taking the time to describe the subtle nuances and detail of the entire process.
Finally here at The Next Platform, we are reminded of more pioneering early days of software and systems development. The current state of R and Deep Learning techniques are similar to early “package management” days. A bewildering array of potential methods are presented to data scientists today. If your problem is selecting the actual package, then it is also clear there are much bigger problems at play. Claims of performance, accuracy and all others have to be immediately set aside. Significantly more research will be needed to go beyond the “hype” curve of all the various and assorted AI technologies and tool kits, especially if (to our main point here) those techniques aren’t documented clearly with example code that is also fully reproducible.
 Dr. James Cuff
Dr. James Cuff
Distinguished Technical Author, The Next Platform
James Cuff brings insight from the world of advanced computing following a twenty-year career in what he calls “practical supercomputing”. James initially supported the amazing teams who annotated multiple genomes at the Wellcome Trust Sanger Institute and the Broad Institute of Harvard and MIT.
Over the last decade, James built a research computing organization from scratch at Harvard. During his tenure, he designed and built a green datacenter, petascale parallel storage, low-latency networks and sophisticated, integrated computing platforms. However, more importantly he built and worked with phenomenal teams of people who supported our world’s most complex and advanced scientific research.
James was most recently the Assistant Dean and Distinguished Engineer for Research Computing at Harvard, and holds a degree in Chemistry from Manchester University and a doctorate in Molecular Biophysics with a focus on neural networks and protein structure prediction from Oxford University.
Follow James Cuff on Twitter or Contact via Email



Be the first to comment